
Mangroves act as valuable nurseries for many coral reef fishes. Protected among the roots and surrounded by nutrient rich water, juvenile coral reef fishes can mature with ample food and protection until they migrate out to join the adult population on offshore reefs. (Illustration E. Paul OBerlander WHOI)
The ecological integrity of coastal habitats is coming under increasing pressure from human activities, such as overfishing and coastal development, as well as a rapidly changing climate. Many fishes are thought to use coastal wetlands, including mangroves and seagrass beds, as juvenile nurseries before migrating offshore to join the adult population. Identifying essential habitats and preserving functional linkages among these habitats is likely necessary to promote ecosystem health and sustainable fisheries. This necessitates quantitative assessment of functional connectivity among essential habitats at the seascape level. This project presents the development and first application of a method for tracking fish migration using amino acid δ13C analysis in otoliths. We have explored the potential for geochemical signatures in otoliths of snapper to act as natural tags of residency in seagrass beds, mangroves and coral reefs in the Red Sea, Caribbean Sea and Eastern Pacific Ocean. The δ13C values of otolith essential amino acids varied as a function of habitat type and provided a better tracer of residence in juvenile nursery habitats than conventional bulk stable isotope analyses (SIA). Using our otolith amino acid isotope fingerprinting approach, we have quantified the relative contribution of coastal wetlands and reef habitats to a common and commercially important meso-predator, the Ehrenberg’s snapper (Lutjanus ehrenbergii), on coastal, shelf and oceanic coral reefs in the Red Sea. We found that juvenile L. ehrenbergii were capable of making amazing long distance migrations from over 30 km from coastal juvenile nurseries to offshore coral reefs, as well as across deep open water that has long been considered to be a hard migration barrier for coral reef fishes. Coastal wetlands were important nurseries for L. ehrenbergii; however, there was significant plasticity in L. ehrenbergii juvenile habitat requirements. Seascape configuration played an important role in determining the functional connectivity of L. ehrenbergii populations in the Red Sea. The compound-specific SIA approach developed in this project will be particularly valuable for tracking the movement of species and life-stages not amenable to conventional tagging techniques. This work provides quantitative scientific support for establishing realistic population connectivity models that can be used to design effective marine reserve networks. This work was established in the Red Sea during my doctoral research in the MIT-WHOI Joint Program in Oceanography, and has since been expanded to other regions and species to gain a more global understanding of the role of nursery habitats to coral reef fishes.
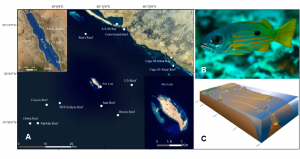

(A) Collection sites from coastal wetlands (Al Lith Bay and Cape Al-Askar Bay), coastal reefs (Coast Guard Reef and Cape Al Askar Reef), shelf reefs (Ron’s Reef, LJ’s Reef, Saut Reef, and Brown Reef), a continental island (Abu Latt), and oceanic reefs (Shi’b Sulaym Reef, Canyon Reef, MarMar Reef, and Dohra Reef) near Al Lith, Saudi Arabia in the Red Sea. (B) Ehrenberg’s snapper (Lutjanus ehrenbergii, Peters 1869) is a commercially important reef-associated snapper species in the Indo-West Pacific. (C) Conceptual diagram of habitat configuration and potential seascape connectivity of L. ehrenbergii in the study area. (Figure from McMahon et al. 2012 PNAS)
Primary collaborators: Dr. Simon Thorrold (WHOI), Dr. Michael Berumen (KAUST)

A snapper otolith holds a wealth of information about where that fish spent its juvenile nursery period. (Photo Tom Kleindinst, WHOI)
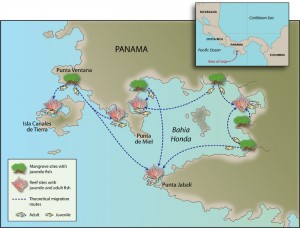
An illustration of essential habitats and migration pathways for coral reef fishes. (Illustration Jack Cook, WHOI)
McMahon KW, Berumen ML, Mateo I, Elsdon TS, Thorrold SR (2011) Carbon isotopes in otolith amino acids identify residency of juvenile snapper (Family: Lutjanidae) in coastal nurseries. Coral Reefs 30:1135-1145 (Best Paper Award finalist)
Popular media:
One man’s swamp is a fish’s nursery. Oceanus Magazine
An ear for history: How a fish’s early life can help determine its future. Nature Middle East
Tracking fish through a coral reef seascape. Woods Hole Oceanographic Institution